I found the answer to my own question. When dealing with millions of data points, you want to avoid letting the front-end doing the color mapping and instead allow datashader to perform this task.
What I struggled with, was getting each curve to plot with separate colors while also displaying a legend for each curve. You can use datashader’s categorical colormapping to handle applying individual colors to each curve. You first have to set the parameter column to type category and then use an aggregator to count by that category:
df["parameter"] = df["parameter"].astype("category")
dask_df = dd.from_pandas(df, npartitions=mp.cpu_count())
dask_df.persist()
datashader = datashade(hv.Curve(dask_df), aggregator=ds.count_cat("parameter")).opts(**options) #
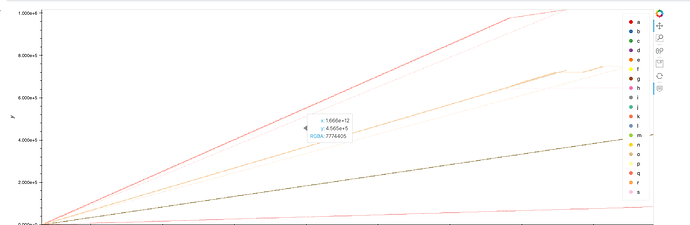
The resulting chart will look something like this: